Overview of the Structure & Function of Hen Egg-White
Lysozyme
Go
back
Lysozyme is a relatively small (129 AA) enzyme that catalyzes the hydrolysis of specific kinds of polysaccharides comprising the cell walls of bacteria.
In birds, lysozyme is an exceptionally abundant protein in fowl egg whites where it likely functions both as an antibiotic (see below) as well as a nutrient for early embryogenesis. In vertebrates, this "glycosidase" is found mainly in biological secretions (such as tears) where it probably serves as an anti-bacterial agent by digesting and weakening the rigid bacterial cell wall, thereby rendering the bacterial susceptible to osmotic lysis. The effect of lysozyme is similar to the effect of penicillin which also weakens the cell walls of bacteria only by irreversibly inhibiting a transpeptidase enzyme required for crosslinking peptidoglycan macromolecules formed in the biosynthesis of the cell wall. Under normal conditions, bacteria grow very rapidly in some cases doubling more than once in an hour. However, when cell wall crosslinking is disrupted, bacteria tend to lyse in hypotonic media as a result of the mechanical weakening of their cell walls.
Lysozyme hydrolyzes a number of structurally similar substrates but the best known substrates for this enzyme are alternating polysaccharide copolymers of N-acetyl glucosamine (NAG) and N-acetyl muramic acid (NAM) which represent the "unit" polysaccharide structure of many bacterial cell walls. Lysozyme cleaves at b(1-4) glycosidic linkage, connecting the C1 carbon of NAM to the C4 carbon of NAG. The optimal substrate is a (NAG-NAM)3 hexasaccharide, with lysozyme cleaving at the NAM4-b-O-NAG5 glycosidic bond. The active site of lysozyme includes binding sites for each sugar ring of the hexasaccharide, these being designated sites "A" through "F." Thus, lysozyme preferentially cleaves at the glycosidic bond connecting ring "D" to ring "E" of the hexasaccharides.

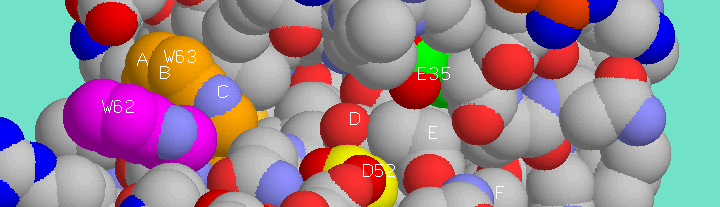
Extended polysaccharide binding site of hen egg white lysozyme. Six sugar monomer binding pockets are indicated, A-F, and the sidechains of two catalytic residues, Glu35 (green) and Asp52 (yellow), are indicated as well as the sidechains of Trp62 (magenta) and Trp63 (orange) which line binding pockets A-C.
Smaller saccharides are effective competitive inhibitors of lysozyme. One such inhibitor, as examined in this web page exercise, is the trisaccharide, NAG3. Tri-NAG binds most efficiently to the saccharide binding sites "A-C" and thus it is positioned away from the key catalytic residues of lysozyme -- namely Glu35 and Asp52 -- which are logically located near the junction between the "D" and "E" binding sites where lysozyme cleaves.
Click on the image below to open the lysozyme web page for further examination of this enzyme's properties.
| © | Duane W. Sears |
| February 24, 2019 |